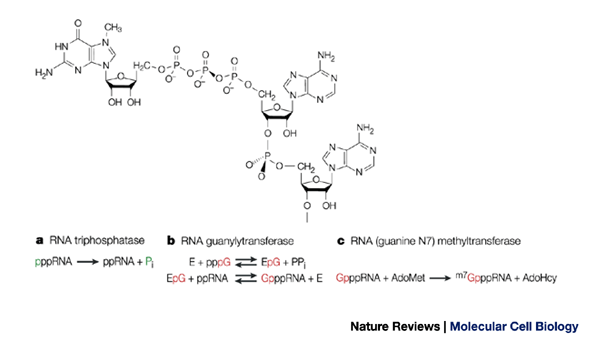
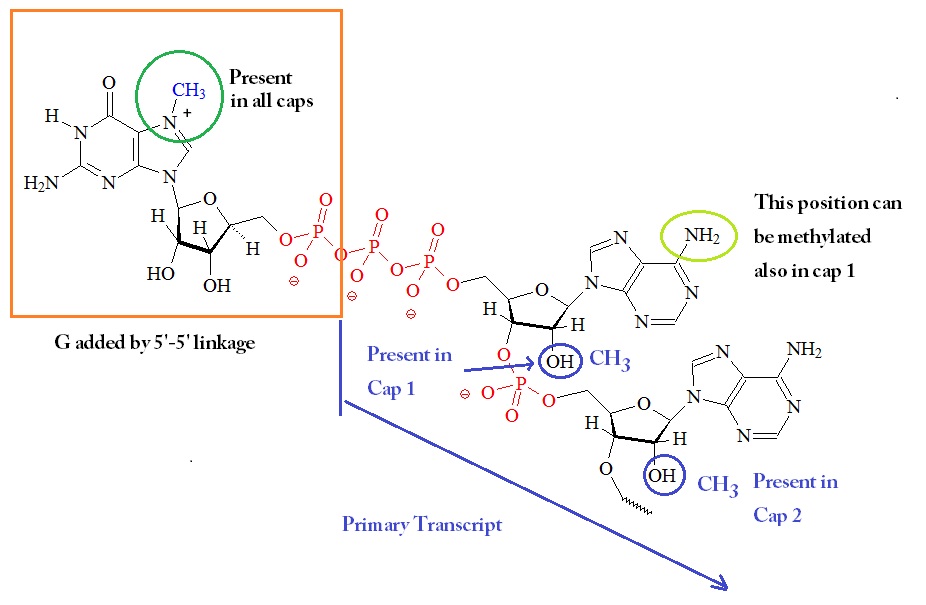
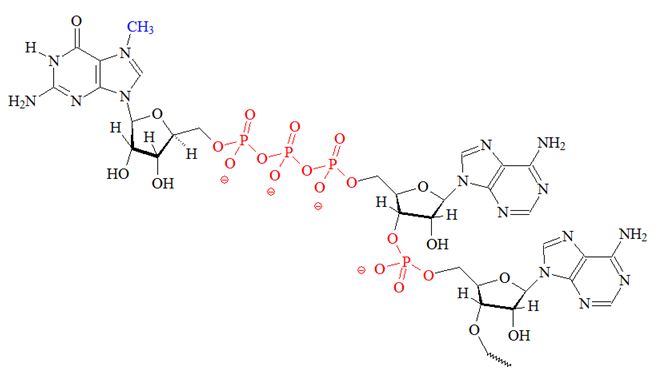
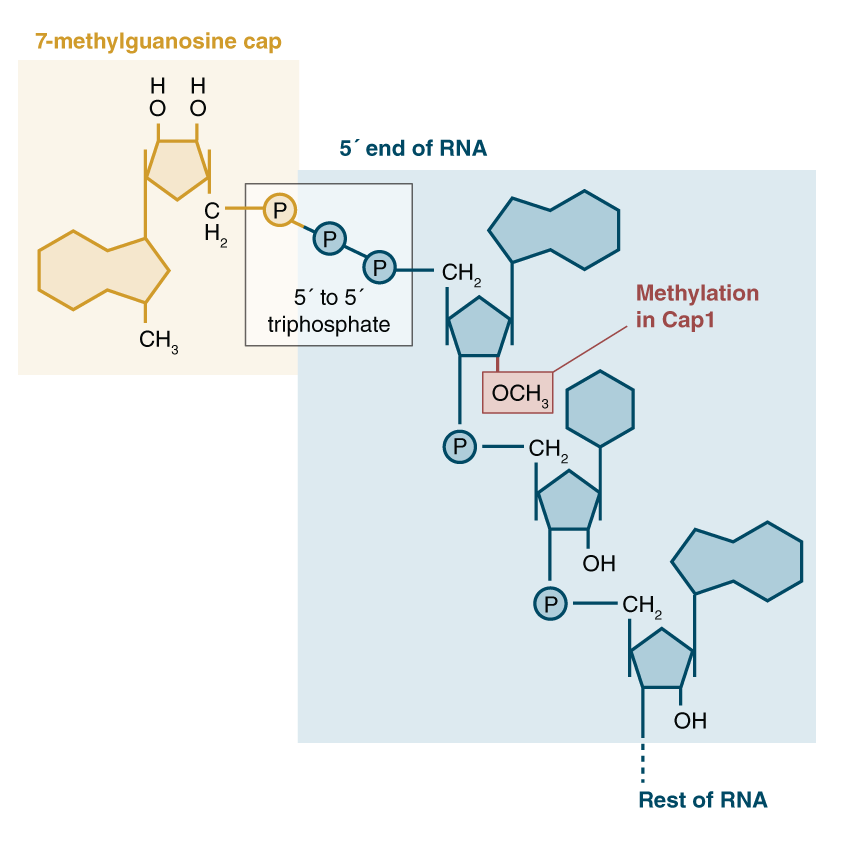
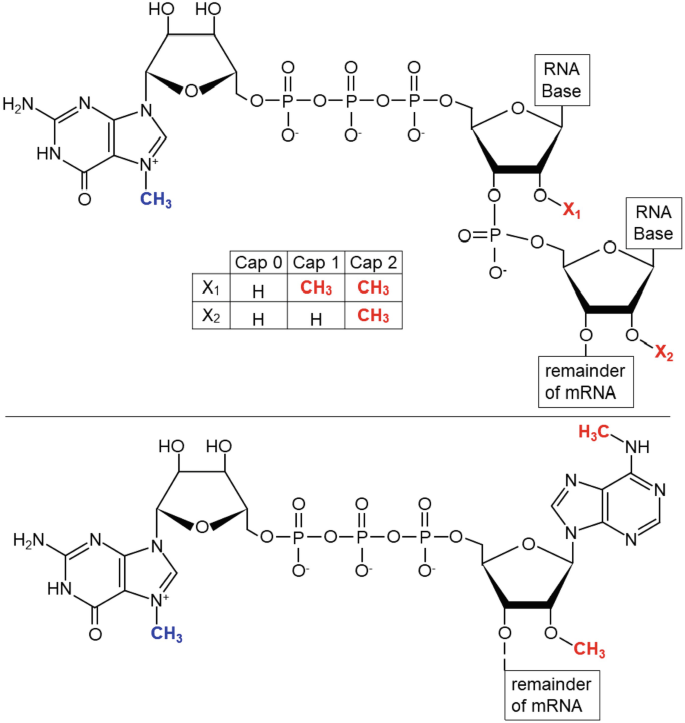
a) The 7-methylguanosine cap. The methyl group of 7-methylguanosine is... | Download Scientific Diagram
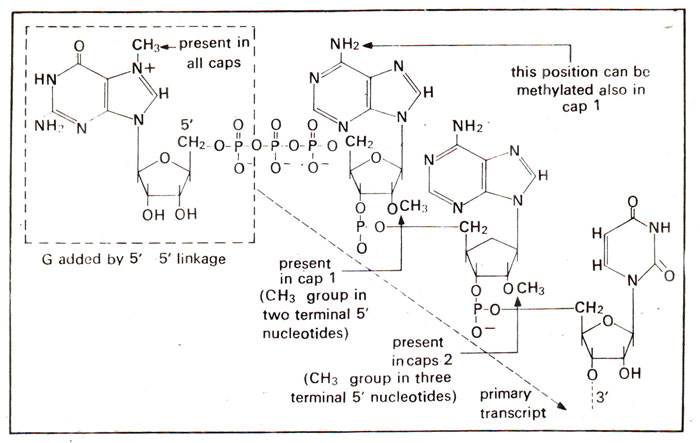
Addition of caps (m7G) and tails (polyA) for mRNA | Expression of Gene : Protein Synthesis RNA Processing (RNA Splicing, RNA Editing and Ribozymes)

The eukaryotic translation initiation factor eIF4E elevates steady-state m7G capping of coding and noncoding transcripts | PNAS

Enzymatic Assays to Explore Viral mRNA Capping Machinery - Kasprzyk - 2021 - ChemBioChem - Wiley Online Library

Cap-specific terminal N6-methylation of RNA by an RNA polymerase II–associated methyltransferase | Science
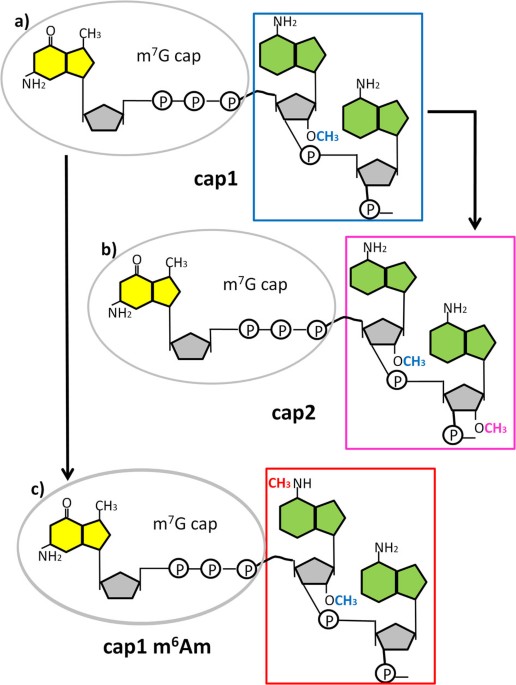
A novel synthesis and detection method for cap-associated adenosine modifications in mouse mRNA | Scientific Reports
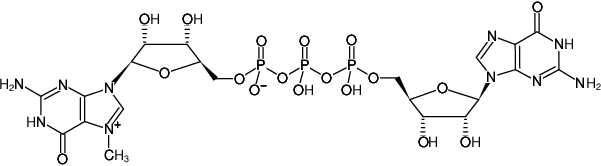
m7GP3G (Monomethylated Cap Analog) - Solution, Cap Analogs – Enhance mRNA stability and translation efficiency - Jena Bioscience

5′-Phosphorothiolate Dinucleotide Cap Analogues: Reagents for Messenger RNA Modification and Potent Small-Molecular Inhibitors of Decapping Enzymes | Journal of the American Chemical Society

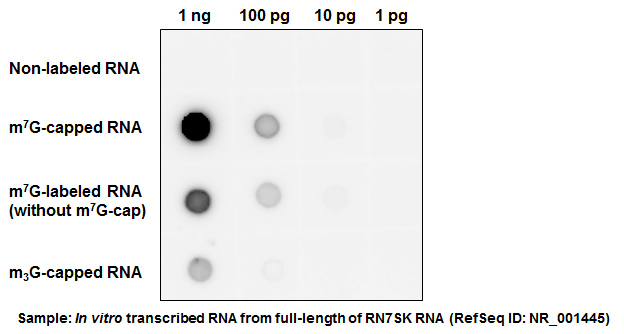
CAP-MAP: cap analysis protocol with minimal analyte processing, a rapid and sensitive approach to analysing mRNA cap structures | Open Biology

5′-Phosphorothiolate Dinucleotide Cap Analogues: Reagents for Messenger RNA Modification and Potent Small-Molecular Inhibitors of Decapping Enzymes | Journal of the American Chemical Society